The Center for Mouse Genome Modification (CMGM) at UConn Health provides design and generation of genetically modified mice and other services.
recombinant adeno-associated viruses (rAAV)
The Center for Mouse Genome Modification (CMGM) is now offering genome editing services using recombinant adeno-associated viruses (rAAV).
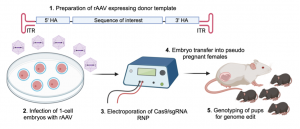
AAV (adeno-associated virus), which is a single-stranded DNA (ssDNA) virus, can effectively penetrate the zona pellucida and infect one-cell embryos. Recombinant AAV (rAAV) contains the sequence of interest together with 5’- and 3’-homology arms for CRISPR-mediated gene editing in the mouse genome. To this end, we infect one-cell embryos with rAAV followed by electroporation of with Cas9/sgRNA ribonucleoprotein (RNP) for subsequent genome modification in the mouse. The advantage of this approach is that rAAV infection is faster and less laborious than traditional pronuclear microinjection to transduce ssDNA template into mouse embryos. Furthermore, ssDNA template in the form of rAAV genome is a more efficient template for homology-directed repair as compared to double-stranded DNA. This approach significantly increases the efficiency to generate novel mouse models containing the sequence of interest.
Piggy Bac mediated transgenesis
New Services
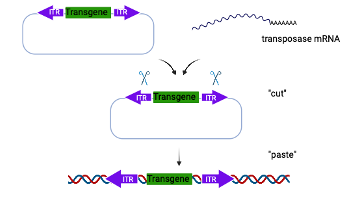
Piggy Bac mediated transgenesis offers many advantages over traditional transgenics. It has a large cargo capacity (up to 200 kb) allowing insertion of large promoters and regulatory elements. The insertion is footprint free and does not leave behind vector sequences. The transgene is also inserted in the host genome in a precise manner as a single copy without any chromosomal rearrangement or deletion (chromothripsis) and therefore offers reliable and consistent transgene expression.
In the Piggy Bac approach, the donor template includes the transgene of interest flanked by inverted terminal repeat (ITR) sequences. The donor template is microinjected into 1-cell embryos along with mRNA encoding the transposase, which integrates the transgene into the host genome through a “cut and paste” mechanism.
The CMGM has generated several novel mouse models using this approach including cell specific ablation and conditional rescue humanized mouse models.
PiggyBac